Research Interests
We work on a number of topics at the interphase between synthetic biology and evolution. We are interested in understanding how evolution shapes the properties of proteins exhibiting quantum mechanisms in vivo, in the tradeoffs governing gene expression and in the dynamics of complex microbial communities. The applications of this research range from the development of novel methods to fight antibiotic resistant bacterial pathogens to the use of engineered microorganisms for the upcycling of plastic waste into molecules of added value.
Experimental evolution
We are interested in developing methods to investigate the evolution of biological information polymers. The complete experimental coverage of a landscape is only possible for very short molecules with a reduced alphabet, such as the one mentioned. In the case of proteins, it is impossible to completely cover the astronomically large space of sequences. It is still feasible, however, to use fitness landscapes to narrow down the search area and find evolutionary paths leading to new functional sequences. We are building on this knowledge to look for bacteriophages and antibodies with evolved specificity for a number of applications, including the development of new biosensors and antibiotics.
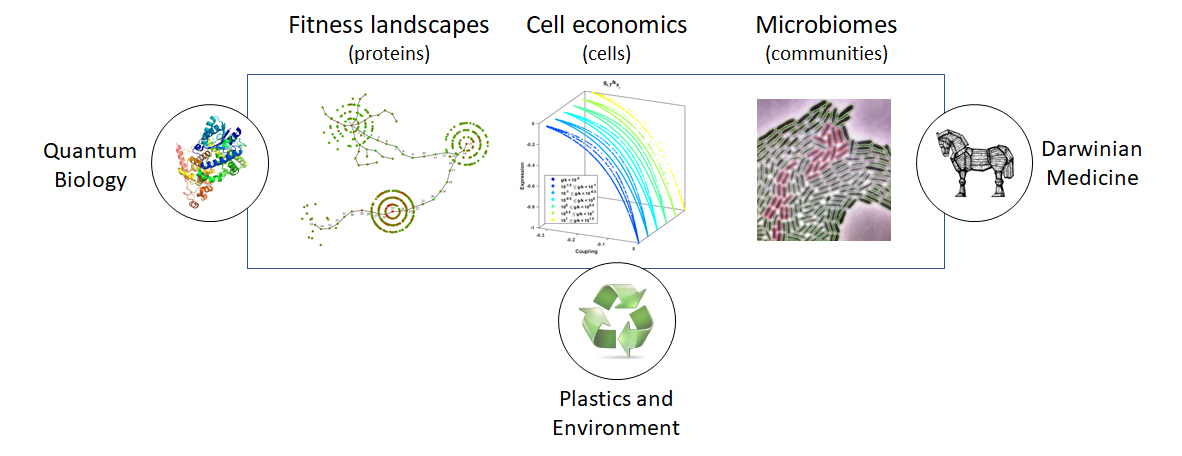
We use fitness landscapes as a metaphor to understand evolution. A fitness landscape is a collection of genotypes performing a particular function. We use the landscape analogy to understand how new functions emerge. For instance, using a combination of selection and high-throughput sequencing we have investigated the ability of a pool of all possible 24mer RNA sequences to bind to a ligand. This pool is composed by 10^14 unique sequences each represented 1,000 times. Besides in vitro experimental evolution we also use in vivo tools, and we are currently leveraging the use of retroelements as tools for lab-directed evolution of genes of interest.
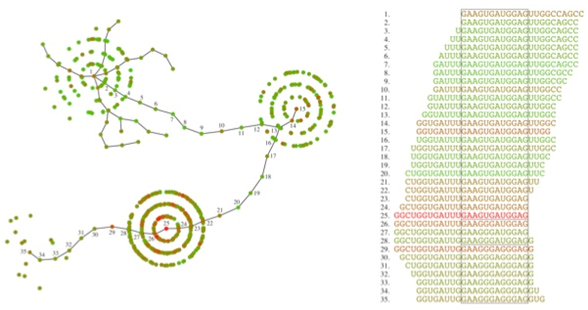
Detail of a two-dimensional representation of a fitness landscape for short RNAs that bing to GTP. The picture shows the evolutionary paths connecting three distinct families clustered in peaks with high activity.
Quantum Biology
Our universe is formed by particles that behave in a quantum and non-classical way. However, at the much larger scale of living cells, which are subject to noise coming from continuous Brownian motion, quantum mechanisms are thought to not to play a relevant role. This view is challenged by some biological processes that are thought to require of quantum properties in order to work efficiently. Among them, we are interested in the magnetic effects present in the blue-light receptor cryptochrome and in the capture of photons carried out by photosynthetic light harvesting complexes. It has been proposed that both of them need of long-lived quantum coherence in order to endow the corresponding proteins with the ability to, respectively, sense the extremely weak magnetic field of the Earth and harvest 98% of the photons received by a leaf.
In these projects we are not only interesting in learning from the quantum mechanisms involved, but we are also trying to assess if they give a selective advantage. In other words, are these processes favoured by evolution so that unlikely quantum properties provide biological systems with a selective advantage? We are also interested in developing applications from this research, focusing on generating biosensors of magnetic fields and more efficient photosynthetic processes in purple proteobacteria.
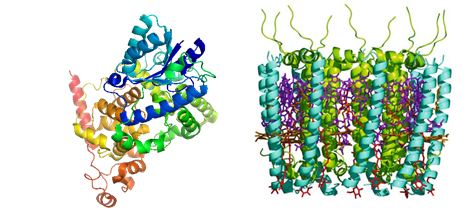
Three-dimensional structures of the proteins cryptochrome (left) and LH2 (right) responsible for sensing magnetic fields and harvesting light respectively.
Cell economics
Cells contain limited amounts of resources available for gene expression. The way these resources are allocated determines cell viability and survival in changing environments. Resource allocation has an impact too in the expression of exogenous genes, typically in synthetic circuits, that generate an extra burden to the cell and fail to behave according to their design specifications. The analysis of how scarce resources are distributed is, in essence, a problem of economics.
We are trying to understand how resources, mainly the transcriptional and translational machinery, are allocated in microbial and mammalian cells. Unravelling molecular determinants of this process helps to optimize the design of synthetic circuits as well as the cellular physiology. We have also developed a strategy to reduce the cost of making proteins based on orthogonal ribosomes that can be used to build feedback controllers. In addition, we are harnessing evolutionary tradeoffs such as the link between bet-hedging functions and growth, in order to optimise the allocation of resources for recombinant gene expression.
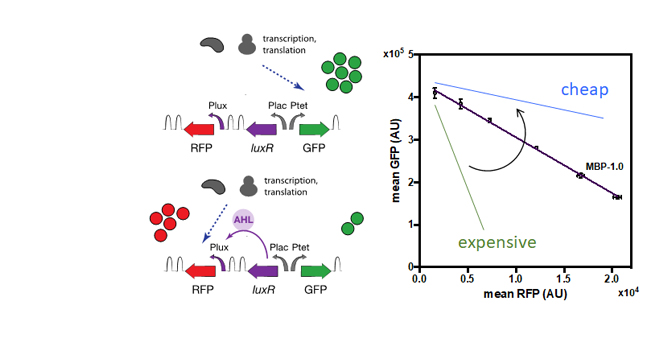
A simple genetic circuit expressing a constitutive GFP and an inducible RFP is used to monitor couplings in the expression of the two proteins due to the competition for RNA polymerase and ribosomes. The couplings are captured by a linear relationship inspired by microeconomics termed ‘isocost line’ that reflects the cellular budget of the cell for making proteins.
Plastics and Environmental Synthetic Biology
We are strong advocates of developing applications of our research. In particular, we have ample experience studying pathways for the removal of either natural or xenobiotic pollutants. Current efforts in this field focus on the degradation of metaloids such as selenite, alkaloids resulting from farming activities and certain types of plastic waste.
Commonly used plastics are relatively new to nature substrates for which microorganisms still have not developed efficient strategies of mineralisation. Using synthetic biology, targeted environmental screenings and lab directed evolution, we are engineering microbial strains for the efficient conversion of non-recyclable plastic waste into molecules with added value. Our lab has recently coordinated coordinating the EU’s ERA-CoBiotec project MIPLACE aimed at integrating PET in the circular economy.
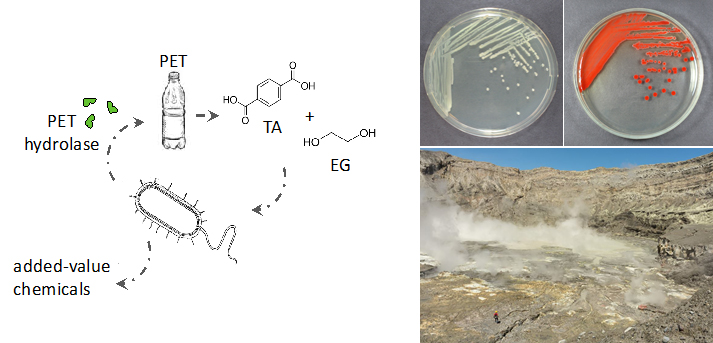
The left panel illustrates the use of PET waste as a feedstock for microbial transformations. We engineer microorganisms to secrete hydrolases that degrade the polymer into the constituent monomers terephthalate and ethylene glycol. The monomers are consumed by the same or other bacteria and they are later transformed into other molecules. The right panel shows the the conversion of naturally occurring selenite present in volcanic areas (picture from sampling trip to the Poas volcano by collaborator Prof. Max Chavarría – Universidad de Costa Rica). Certain species of Pseudomonas can reduce selenite into bright red elemental selenium.
Population dynamics in microbiomes and AMR transmission
Microbes do not live in isolation. Interactions in communities are of great importance for environmental processes and also affect – and are affected by – our lifestyle as exemplified by the spread of antimicrobial resistance genes (AMR).
The transmission of AMR depends on many environmental factors including the presence of large bacterial aggregates called biofilms. In complex populations, limited cell mobility together with limited diffusion of nutrients and other shared molecules, have as a consequence the formation of distinct habitats with differential species composition. The evolution that microbes undergo in these niches is greatly conditioned by surrounding species and their capability to adapt, giving rise to even more complex dynamics and interactions, which condition the efficacy of antibiotic treatments among others.
We are interested in analysing the influence of the environment in microbial social interactions including AMR. To this end we use a combination of lab controlled experiments with simple populations and Next-Gen sequencing in natural communities to investigate the transmission of AMR encoding genes.
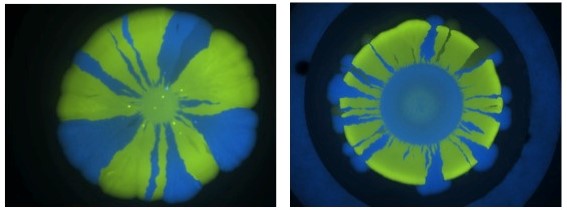
Effect of genetic drift on conjugation on agar plates. Two populations of cells tagged with different colours are allowed to demix and form genotypic identical sectors from the centre of a plate. If one of these populations carries a transferable plasmid it is possible to quantify conjugation challenging the colony with a circular filter soaked in an antibiotic (right picture)
Darwinian Medicine
Darwinian medicine refers to the notion of using biological entities to fight disease. Darwinian medicine advocates for keeping a healthy balance of the microbiome composition so that pathogenic organisms can be kept in check by other members of the community. Examples of this include the use of bacteriophages to fight infectious diseases. In addition to phage therapy, we are exploiting the social interactions that may take place between the therapeutic agents and other biological elements responsible for producing the disease in the first place. Many pathogens require coordinated efforts to colonise a niche (a patient). For instance, Pseudomonas aeruginosa infects patients suffering from cystic fibrosis using sophisticated social interactions such as the secretion of the siderophore pyoverdine.
We have studied the costs and benefits of making pyoverdine in different environmental conditions and we have come up with a strategy, based on the altered expression of pyoverdine receptors, that allows mutant strains to invade communities of P. aeruginosa that are resistant to antibiotics. This strategy could be used to deploy Trojan horses in the community, eventually replacing a bacterial population resistant to an antibiotic with another that is sensitive to it.
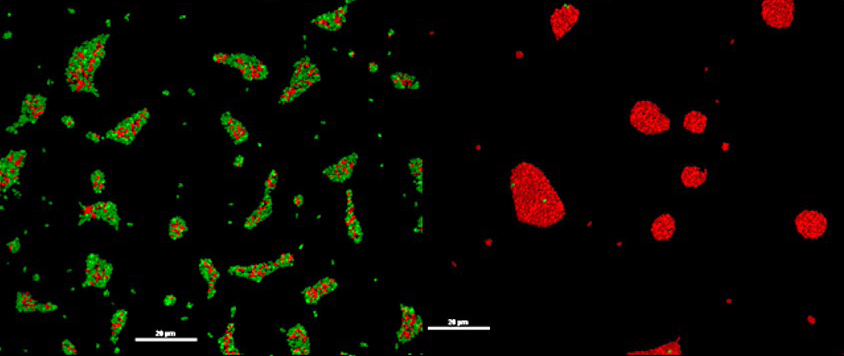
When two strains of P. aeruginosa are co-cultured they compete for the same niche. Under standard growth conditions (left) the wildtype (green) has a selective advantage and colonises the population. However, if the cost of making a public good such as pyoverdine is sufficiently high (right), the population can easily be taken over by a mutant (red) that does not pay as much of the cost and maximises its benefits.
